Discovery
|
Ken Henderson, PhD
About Those Mutant Virus Strains….
The SARS-CoV-2 virus that causes COVID appears to be getting smarter, but can a virus learn?
The idea that a virus particle made up of several proteins, a genome of approximately 30 thousand nucleotides, and a lipid envelope layer that it stole from the host cell, can learn is not far from the truth. Certainly, viruses don’t have the neural capability to figure things out, but they do have natural algorithms based on four possible nucleotide base outcomes in its genetic code.
No different than any other higher evolved organism, viruses follow the same dogma of molecular biology, which is that a three nucleotide base pair sequence or “codon” made up of any combination of four different nucleotides codes for an amino acid. These amino acids are then linked together by ribosomes in the cell cytoplasm to build proteins used for both functions (enzymes) and structures. The human genome consists of a double-stranded deoxyribose nucleic acid (DNA) genome made up of four deoxyribose nucleotides adenine (A), cytosine (C), guanine (G), and thymidine (T). However, the genetic blueprint for SARS-Cov-2 (SARS-2) is a single-stranded ribose nucleic acid (RNA) genome made up of the four ribonucleotides, A, C, G and uracil (U, instead of T). The integrity of the SARS-2 RNA genome, made up of roughly 30 thousand RNA nucleotides, is governed by the virus’ RNA-dependent RNA polymerase (RdRP), which is responsible for synthesizing new copies of the virus RNA genome. The new RNA genome is responsible for a next generation of virus and is loaded up in the newly created virus particle shipped out of the infected cell to the next susceptible cell in the same host or shed to infect a new host.
Where do all the mutations come from?
By no intended mistake, and to the virus’ advantage, the virus RdRP is sloppy. Although in humans, our nucleic acid polymerases are more accurate when copying our genomic nucleic acid, RdRP in viruses has a notoriously high rate of single nucleic acid base mistakes called point mutations. So instead of inserting a “C”, it may insert a “U,” “A” or a “G.” Not all point mutations will change the encoding of an amino acid, since often a mutation in the third base of a codon will code for the same amino acid. Conversely, a mutation, in the first two nucleotides of a codon will likely lead to a change in the coded amino acid incorporated in a protein. Less often, one or more nucleotides may be deleted or inserted into the genome by the polymerase, which may cause a ripple effect in all the three base pair codons downstream.
Some viruses such as influenza have multiple segmented genomes so that whole segments can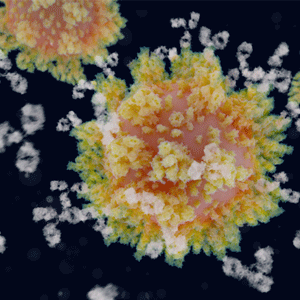 be swapped with other influenza viruses that simultaneously infect the same cell. A change in one or more amino acid in a protein will change the shape and function or function efficiency of the expressed protein. However, whether the new mutations in the virus genome will be an advantage or compromise the virus, will not be determined until the virus containing the modified virus genome enters a new cell. There, it will attempt to use its modified genome to hijack the cell’s machinery for the purpose of producing thousands of copies of itself, which in turn will also carry new mutations.
be swapped with other influenza viruses that simultaneously infect the same cell. A change in one or more amino acid in a protein will change the shape and function or function efficiency of the expressed protein. However, whether the new mutations in the virus genome will be an advantage or compromise the virus, will not be determined until the virus containing the modified virus genome enters a new cell. There, it will attempt to use its modified genome to hijack the cell’s machinery for the purpose of producing thousands of copies of itself, which in turn will also carry new mutations.
Mutations are passed on from one generation to the next. The longer the lifecycle, and the fewer the offspring the slower mutations will occur and be passed on to the next generation. Humans after reaching childbearing years can reproduce every 9 months, and after 3 generations could theoretically have well over a hundred offspring within 60-70 years. Mice can begin breeding at 4-7 weeks of age and could have over 2,500 offspring in less than 6 months! SARS-2 within one cell can produce 1,000 virions in less than a day, and after a week, the infected host can produce over 10 11 virions or 100 billion virons. The short life cycle of a virus and potential for high replication expedites the introduction and diversity of virus mutations.
As the RdRP is incorporating nucleotides into a newly synthesized virus genome, each nucleotide presents the possibility for inserting a mutation within the SARS-2 RNA genome, but a high percentage of virus mutations are never passed on to the next generation because the mutation is fatal to the virus. For example, an amino acid change in the portion of the RNA genome that encodes the RdRP is not likely to be passed on to progeny virus because the precise enzymatic shape and function of the RdRP is required to synthesize the next generation of viruses. That is why genes encoding functional proteins responsible for virus RNA or assist in protein synthesis tend to be conserved (fewer mutations) in a virus genome.
However, amino acid changes that modify structural proteins which are exposed to the host’s immune system on the outside of the virus can potentially serve as a mask that is not recognized as a previously invading strain. A mutation in a structural protein usually does not impact enzymatic function and it is usually an advantage, unless alteration prevents the virus capsids from correctly forming or grossly modifies a binding site that prevents entry into a host cell. Genes that encode structural proteins usually also have the greatest amount of nucleotide variation because they are favored by evolution to evade the host’s immune system.
The SARS-CoV-2 spike protein
So in what ways can mutations in the SARS-2 genome improve virus transmission or appear to cause more severe disease? Changes in the SARS-CoV-2 virus spike protein, which is responsible for attachment and entry into target cells via an angiotensin-converting enzyme 2 (ACE-2) receptor, can lead to a virus being more infectious. Some reports suggest that one point mutation resulting in an amino acid change has changed the shape of the region surrounding the ACE-2 binding site on the spike protein, which makes the virus binding site more exposed and improves the chance for interacting with the host cell’s receptor. Another mutation that can slightly change the shape of the spike protein could also improve the affinity of the virus binding site to the ACE-2 receptor. A titer bond would allow the virus to bind more efficiently to the ACE-2 receptor site and increase cell infection rates.
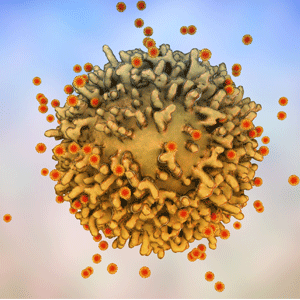 The potential to improve the binding capability of the virus to the host cell is one way to augment infection. Another way is to improve the efficiency of virus replication. The more infectious virus particles a cell can be exploited to produce, the higher amount of virus is shed by the infected person, which can go on to infect others. It only takes one functional virus to find a receptor in a new host to start an infection; therefore, the more virus you are exposed to, the greater the possibility that an infection event will occur. Viruses for the most part, hijack cells and take over their machinery in order to do one thing, make more virus. Any modifications of the virus genome that can improve viral RNA or virus protein synthesis efficiency can lead to an increase in the number of functional virus particles that are released from that cell to infect elsewhere. One study showed that SARS-CoV-2 has a higher reproductive number compared to SARS-1 (2003 outbreak), which is one factor that is believed to have contributed to the success of SARS-2 transmission versus SARS-1.
The potential to improve the binding capability of the virus to the host cell is one way to augment infection. Another way is to improve the efficiency of virus replication. The more infectious virus particles a cell can be exploited to produce, the higher amount of virus is shed by the infected person, which can go on to infect others. It only takes one functional virus to find a receptor in a new host to start an infection; therefore, the more virus you are exposed to, the greater the possibility that an infection event will occur. Viruses for the most part, hijack cells and take over their machinery in order to do one thing, make more virus. Any modifications of the virus genome that can improve viral RNA or virus protein synthesis efficiency can lead to an increase in the number of functional virus particles that are released from that cell to infect elsewhere. One study showed that SARS-CoV-2 has a higher reproductive number compared to SARS-1 (2003 outbreak), which is one factor that is believed to have contributed to the success of SARS-2 transmission versus SARS-1.
Viral factors driving morbidity and mortality
How can a mutation in the SAR-2 virus lead to more severe disease? As a result of exploiting the cell’s resources to make more virus, a SARS-2 infected cell will eventually die. Cell death is often one of the key events that activates the host’s immune system to fight the virus. The death of some cell types are more noted by the immune system than others because of the cell-specific biomolecules a particular cell type might release during cell death (apoptosis). Some classes of biomolecules may serve to elicit the immune system to the site of cell death and contribute to inflammation. Virus-infected cells will eventually go through a programed apoptosis to naturally curtail the virus replication.
Responding immune cells also eliminate the infected cells and prevent more virus from being produced. However well-evolved viruses can actually hold-off apoptosis in favor of continued virus production or to maintain stealthiness within the host cell. SARS-2 usually reaches a rapid high virus count peak in the lungs within the first week compared to SARS-1 which can take up up to 2 weeks to peak and at a lower virus titer. As a result, the immune response to SARS-2 is more abrupt and elevated, and eventually its host’s own immune anti-inflammatory response can lead to pathogenesis in the lungs and resulting death. Although the ACE-2 receptor is predominantly in the respiratory tract, the receptor and resulting infection can also be found in cells within the intestinal tract and epithelium of the kidneys, blood vessels, and heart which can explain observed gastrointestinal and cardiovascular complications. Certainly, both the broad availability of the ACE-2 receptor across different organs and being a high-number propagating virus has defined SARS-2, but any continued mutation that will improve virus receptor binding or the replication number will have an impact on the severity of pathogenesis.
produced. However well-evolved viruses can actually hold-off apoptosis in favor of continued virus production or to maintain stealthiness within the host cell. SARS-2 usually reaches a rapid high virus count peak in the lungs within the first week compared to SARS-1 which can take up up to 2 weeks to peak and at a lower virus titer. As a result, the immune response to SARS-2 is more abrupt and elevated, and eventually its host’s own immune anti-inflammatory response can lead to pathogenesis in the lungs and resulting death. Although the ACE-2 receptor is predominantly in the respiratory tract, the receptor and resulting infection can also be found in cells within the intestinal tract and epithelium of the kidneys, blood vessels, and heart which can explain observed gastrointestinal and cardiovascular complications. Certainly, both the broad availability of the ACE-2 receptor across different organs and being a high-number propagating virus has defined SARS-2, but any continued mutation that will improve virus receptor binding or the replication number will have an impact on the severity of pathogenesis.
Immune escape
How do mutations help the virus evade our host’s immune system? As the SARS-2 pandemic progressed across the globe, the number of people infected with the virus and eventually recovered is now well beyond 100 million people. Recovering individuals have gained a trained immune system to protect from reinfection with the same initial strain, with one study reporting >90% of study participants having long-term immunity beyond 5 months post-exposure. During SARS-2 infection, a subset of the T- and B-cell lymphocytes, capable of recognizing the virus proteins, work together to produce an antibody response specific for SARS-2. Some of these antibodies can directly neutralize the virus and prevent infection of future cells by binding and blocking the binding site for the ACE-2 receptor.
Other non-neutralizing antibodies that bind to the virus can be recognized by macrophages which can consume the antibody bound virus. Although over time, protective antibody levels drop post-infection , subsets of T and B lymphocytes, referred to as memory cells, can remain for years after an infection and become reactivated by the presence of the virus antigen so that there is a more rapid immune response with a second infection.
This seems like a fool-proof way to ward off future infection of the same virus strain, but viruses can adapt. Nucleotide mutations leading to amino acid changes in the virus structural proteins can prevent neutralizing antibodies your body has created for one strain from protecting you from a new strain. Similarly, treatments such as monoclonal antibodies (made up of only one cloned antibody) that target a specific SARS-2 strain may be unable to bind to its neutralizing target if a mutation alters the shape of their binding site. Even if neutralizing epitopes are not effectively bound for new mutant virus strains, there still may be partial binding that will reduce the impact of a second infection. Luckily, your body has a library of B and T cells capable of targeting foreign proteins not recognized as “self” and your body can redirect its targeting antibody efforts towards the new virus strain.
Virus mutation is an evolutionary process where those viruses that can enter a cell more easily or replicate to higher numbers will quickly outpace other viruses lacking those advantages and become the dominant strain. Certainly, the emergence of new mutant strains that may have already compromised the effectiveness of current vaccines is concerning. Host behavior is an important and powerful influence for the creation and continued existence of new virus strains. Viruses that infect individuals that self-isolate to prevent transmission to others will promote a “dead-end” scenario for the new virus mutants produced in that host because those viruses can no longer contribute to the virus gene pool. A virus with a mutation must replicate in a new host for that mutation to be multiplied and passed on to others.
replicate to higher numbers will quickly outpace other viruses lacking those advantages and become the dominant strain. Certainly, the emergence of new mutant strains that may have already compromised the effectiveness of current vaccines is concerning. Host behavior is an important and powerful influence for the creation and continued existence of new virus strains. Viruses that infect individuals that self-isolate to prevent transmission to others will promote a “dead-end” scenario for the new virus mutants produced in that host because those viruses can no longer contribute to the virus gene pool. A virus with a mutation must replicate in a new host for that mutation to be multiplied and passed on to others.
On the other hand, an active social behavior permits a virus with a mutation to continue to exist and replicate among contacts, and a disregard for protection measures will only disseminate the virus more quickly among new contacts. It is in this environment that those individuals who had previously been infected with one virus strain are then more likely to be exposed to and replicate new mutant viruses that are not recognized by their immune system. These individuals can now serve as hosts for replication and transmission of mutant strains that can continue to take advantage of those that are more socially active.
Lastly, it is not the intention of a virus to kill the host since once that occurs the virus is not passed on to future hosts and therefore cannot survive. A successful virus will mutate and evolve so that it learns to live with the host over many generations. There are viruses that have evolved with humans which are flowing through your blood this very minute. For the most part, many of these viruses are harmless and may play an important role in keeping our immune system on alert. Most reading this article have also likely been infected, through other people, with coronaviruses related to SARS-2 (beta-coronaviruses). These coronaviruses have adapted and evolved with us to only cause a “mild cold”, but it is believed that even these viruses likely began in a bat a long time ago just as SARS likely did.

