Industry Focus
|
Melvin Lye, Christoph Eberle, PhD
AI in Flow Cytometry: Hype, Friction, and an Unavoidable Shift
Artificial intelligence (AI) tools deployed for flow cytometry are no longer experimental, but operational
AI systems now perform automated quality control, detect acquisition instability, classify immune sub sets, and in some cases predict clinical outcomes directly from high-dimensional data. Platforms such as DeepCyTOF, CyGate, FlowSOM, Phenograph, and emerging deep neural architectures demonstrated that supervised and unsupervised learning can match or exceed expert gating performance under controlled conditions. Clinical workflows assisted by AI have reported strong concordance with hematopathologist-derived subset percentages and dramatically reducing turnaround time. From the outside, the trajectory appears clear. AI works.
sets, and in some cases predict clinical outcomes directly from high-dimensional data. Platforms such as DeepCyTOF, CyGate, FlowSOM, Phenograph, and emerging deep neural architectures demonstrated that supervised and unsupervised learning can match or exceed expert gating performance under controlled conditions. Clinical workflows assisted by AI have reported strong concordance with hematopathologist-derived subset percentages and dramatically reducing turnaround time. From the outside, the trajectory appears clear. AI works.
Adoption should accelerate. Yet, across CROs, biotech laboratories, and clinical centers, AI remains unevenly deployed. Many labs experiment with it; few rely on it as infrastructure. The reason for this surfaced during recent discussions at the NIST Flow Cytometry Consortium. What emerged was not skepticism about AI capability. It was something more fundamental: a recognition that AI exposes structural weaknesses in how we generate, standardize, and govern cytometry data. The friction is not computational. It is infrastructural.
Instrument Reality AI Cannot Ignore
In the consortium update on CAR-T interlaboratory characterization, one statement cut through the technical details: “The cytometer landscape is still very diverse”. BD Canto systems remain in regulated environments. Lyric platforms are replacing older hardware. CytoFLEX instruments coexist with spectral Aurora systems. MACSQuant analyzers participate alongside legacy configurations. Panel substitutions were required because not all instruments possessed identical detector architecture. In some cases, fluorophores had to be replaced simply because specific violet channels were unavailable.
This is not an inconvenience. This is structural heterogeneity. AI models trained on a single signal representation implicitly assume the detector geometry, excitation energy, spillover structure, and compensation behavior. When deployed onto a different instrument family, they encounter distributional shift. What manual gating tolerates, neural networks amplify. The conversation around tandem dyes illustrated this vividly. A participant cautioned that tandem emission spectra can vary depending on excitation power and beam geometry, potentially leading to inconsistencies across cytometers. This is not a new concern. But it becomes consequential when models are sensitive to subtle intensity distributions. Manual gating tolerates subtle distribution shifts. Deep learning does not. The reliability of AI in flow cytometry depends on the model architecture and the quality of the input data. While AI models can produce unreliable results when used blindly, they generally provide laboratories greater consistency than manual gating.
Compensation As Epistemological Question
The discussion of compensation matrices went beyond procedural detail. Matrix selection was evaluated based on spillover spread metrics and performance across instruments. Some platforms favored cell-based autofluorescence extraction; others relied on bead controls. Iterative approaches, such as auto-spill, were considered within the context of holistic matrix optimization. What this conversation revealed is that compensation is not merely a technical correction. It is epistemology. It encodes assumptions about how fluorescence should behave. When AI models are trained downstream of compensation, they inherit those assumptions. Two laboratories may generate biologically identical samples yet feed computational systems measurably different signal distributions. AI cannot adjudicate which compensation philosophy is correct. It simply reflects the consequences.
Confront foundational preprocessing
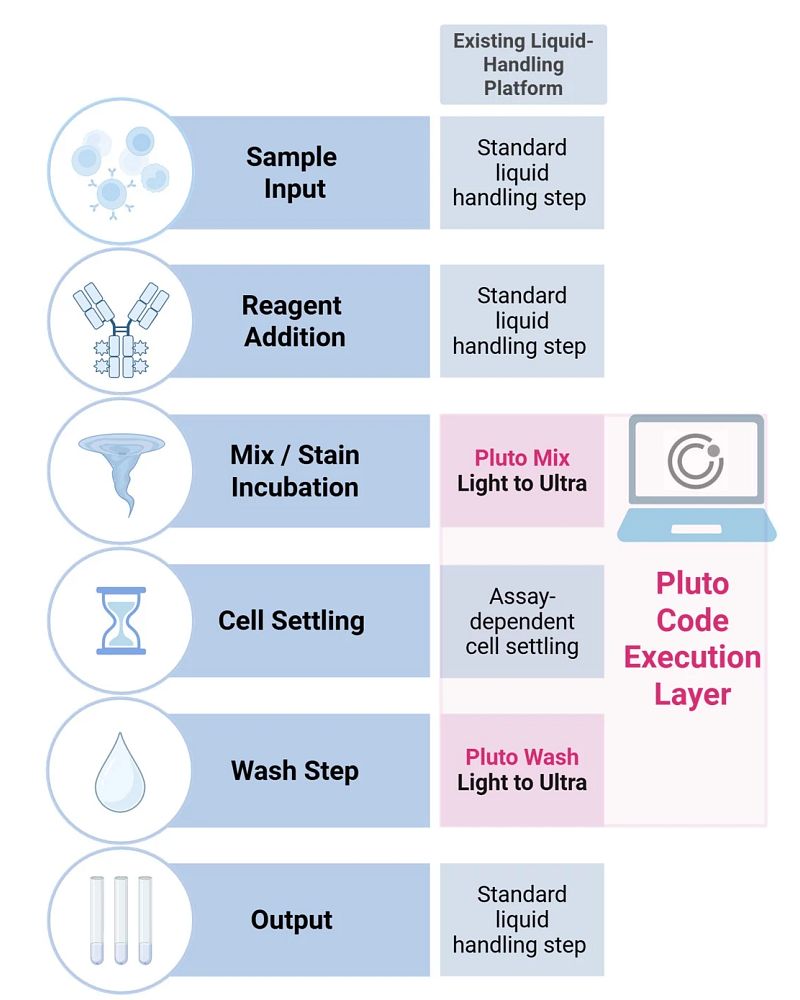
Figure 1. Pluto Code-enabled execution of C-FREE centrifuge-free washing within standard liquid-handling workflows. Pluto Code implements C-FREE gentle wash behavior as executable Pluto Mix and Pluto Wash primitives that run natively within a liquid handler’s method editor. User-authored steps (sample input, reagent addition, output) are combined with code-defined mixing, assay-dependent cell settling, and tunable wash stringency (Light to Ultra), enabling centrifuge-free, on-deck sample preparation while preserving workflow structure across liquid-handling platforms.
Rare Populations: Promise and Statistical Fragility
The debate over event counts was telling. One participant referenced the informal “10,000-event standard.” Others noted that, depending on the biological context, far fewer cells may suffice for frequency estimation. In classical manual gating, abundant populations remain robust even with moderate sampling. But AI systems trained to detect rare or phenotypically shifting populations (tumor-infiltrating lymphocytes, CAR-T subsets, activated memory compartments) depend on both representation richness and statistical stability. Supervised frameworks, such as CyGate, demonstrate that rare subsets can be reliably identified when supported by sufficiently rich expert-labeled datasets. However, the ceiling of performance is constrained by label quality and diversity of representations. Inter-expert variability propagates into model bias.
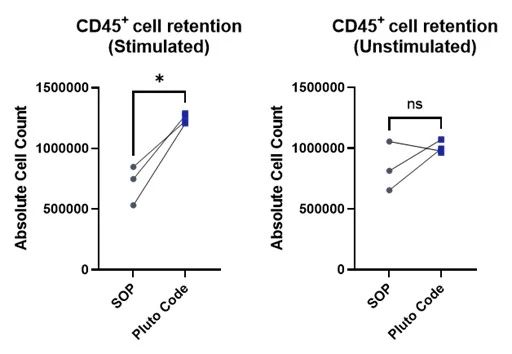
Figure 2. CD45+ cell retention of stimulated and unstimulated PBMCs prepared by Pluto Code or centrifuge-based SOP on an Opentrons OT-2 platform. Statistical analysis by paired Student’s t-test.
In rare-event detection, AI’s strengths become its vulnerabilities. It excels at high-dimensional discrimination but struggles when the underlying measurement noise exceeds the biological signal. The solution is not “better models.” It is a better measurement discipline.
From Automated Gating to Representation Learning
The most forward-looking portion of the consortium discussion did not center on autogating tools. It focused on representation learning, embedding, and latent space modeling. This marks a conceptual shift. Each cytometry event is not simply a vector of marker intensities. It is the product of instrument architecture, detector calibration, fluorochrome chemistry, SOP versioning, reagent lot, sample preparation, compensation approach, and biological context. These variables constitute a multi-layered feature space far richer than the typical 20–40 measured channels. Embedding approaches (autoencoders, contrastive learning, transformer-based architectures) offer a path toward mapping such structured data into latent representations that may reduce platform bias. The question posed implicitly was profound: can T cells measured on Aurora cluster in latent space with T cells measured on CytoFLEX despite hardware differences? If yes, AI becomes cross-platform robust. If not, it remains instrument bound. The future of cytometry AI depends on that answer.
The Illusion of Generalization
Most readers would agree that AI performance often degrades when encountering new instruments, panels, fluorochrome combinations, or sample types not represented in training data. Deep architecture can reduce technical variation across studies, but residual batch effects persist. Many published results demonstrate impressive AUROC metrics across curated datasets. However, cross-study benchmarking remains inconsistent due to differences in preprocessing, annotation, and metrics.
Generalization claims in cytometry AI must therefore be interpreted cautiously. Without standardized reference datasets and transparent benchmarking protocols, performance comparisons lack structural grounding. The consortium’s emphasis on centralized data analysis pipelines, standardized file naming, QC review, and AI-ready dataset generation reflects an emerging recognition: interoperability must precede scalability.
The Structural Shift
We are moving through three phases of AI maturity for cytometric applications:
- The first phase (automated gating and QC) is operational today. Tools such as DeepFlow™ and CyGate demonstrate that supervised learning can replicate expert-defined strategies in controlled settings.
- The second phase (cross-platform representation learning) is emerging. Here, the focus shifts from reproducing manual gates to harmonizing signal spaces across instruments.
- The third phase (a foundation-style cytometry model that is panel-aware, platform-aware, and transferable across institutions) remains aspirational.
Achieving it will require more than computational innovation; it will also require disciplined SOP alignment, reagent stability, detector calibration, and shared ontological frameworks. AI will not standardize cytometry for us. Users must standardize cytometry for AI.
The Strategic Inflection Point
The consortium discussions were not about hype. They were about discipline. File naming conventions were standardized. Compensation matrices were scrutinized. Viral homogeneity was quantified. SOP revisions were debated with precision. This is the quiet work that enables AI maturity. The most dangerous misconception in our field today is that AI adoption is primarily a software decision.
It is not. It is a structural commitment to measurement consistency, metadata integrity, and cross-platform comparability. The future of AI in flow cytometry will belong to laboratories that treat data generation as infrastructure rather than output. No longer is the question whether AI could analyze flow cytometry data, but: “Can we build a cytometry ecosystem suitable for AI?”
Bench + Bytes is a column written by Charles River Scientist Christoph Eberle, PhD, and Melvin Lye, Senior Director, Scientific Affairs and Product at Curiox Biosystems. It is hosted by Eureka, Charles River's scientific blog.