Microbial Solutions
|
Ryan Cox, BS
An Emerging Pathogen of Interest
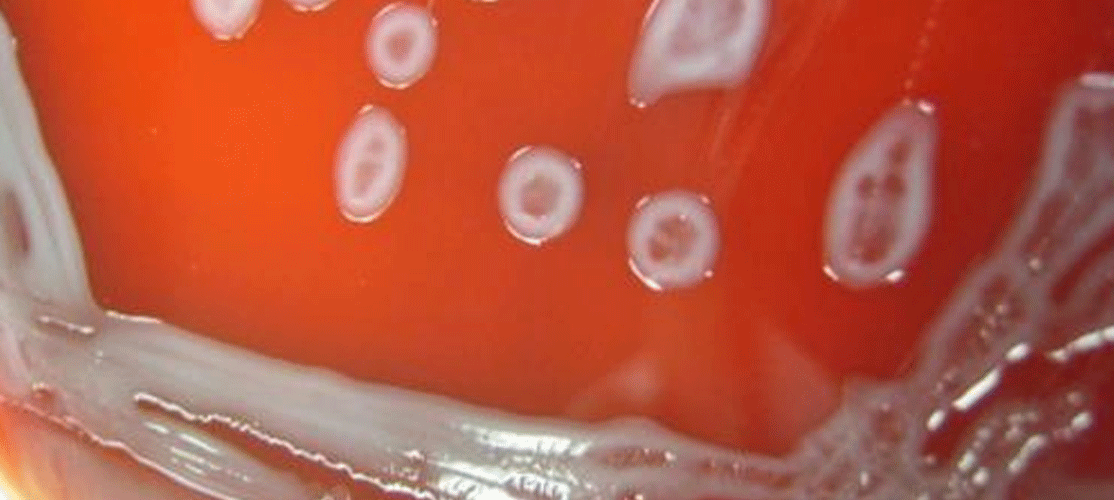
The bacterium Leclercia adecarboxylata, which the FDA is watching, has been linked to several hospital outbreaks
Following reports of Leclercia adecarboxylata contamination of platelet (PLT) components , the US Food and Drug Administration (FDA) increased its scrutiny of blood product collection facilities, reinforcing the organism's link to catheter-related infections and proclivity to multiply in blood [1] [2]. While these contamination events are rare, the FDA's recommendations to blood establishments and transfusion services demonstrate the prevalence of an emerging pathogen that has previously been hidden in the shadows.
Early ID and strain typing of bacteria help determine treatments
Modern microbial identification methods have been critical in identifying L. adecarboxylata as an emerging pathogen of interest. L. adecarboxylata was recently identified as a common organism isolated from polymicrobial infections in immunocompetent patients [3]. Several other publications discussed a much more serious situation caused by product contamination, which resulted in an outbreak. These L. adecarboxylata hospital outbreaks were caused by contaminated heparin vials while another case was caused by using contaminated total parenteral nutrition (TPN). Both early identification and strain assessment play important roles in giving patients proper treatments and eradicating product contamination as soon as possible. In recent years, cases of reported Leclercia infection have increased in frequency. The taxonomic conundrum surrounding the organism has established the need for more specific methods of identification.
L. adecarboxylata belongs to the Enterobacteriaceae family. This family is the largest in the order Enterobacterales consisting of 28 genera and over 75 validly published species, including many pathogens such as Escherichia coli, Klebsiella pneumoniae, and Salmonella enterica. Because of their diverse properties and historical role in studies focusing on genetic exchange, biochemical pathway elucidation, genetic map sequencing, gene regulation, genetic engineering, and molecular portrayal of viral morphogenesis, members of this family, particularly Escherichia coli and Salmonella, occupy a central position in current biology [4].
Until recently, the order Enterobacterales consisted of only one family, Enterobacteriaceae, and many of the genera within this order are misclassified and polyphyletic (containing multiple species derived from independent origins) [5]. Initially, many species in these genera were classified based on clinical descriptions rather than genotypic characterization. Although reclassification attempts based on single and multiple housekeeping genes have been made, the homogeneity within the Enterobacterales order makes it difficult to reliably determine species using 16S rRNA genes. As a result, many organisms with similar genotypic resolution that should have been classified as belonging to the same genus are instead classified as belonging to different genera. The genus Leclercia is an example of this.
Until recently, Leclercia was a monotypic genus represented by Leclercia adecarboxylata alone. After identifying a group of yellow pigmented organisms that resembled Escherichia coli using the IMViC test (a test for identifying members of the family Enterobacteriaceae), Henri Leclerc proposed the nameEscherichia adecarboxylata in 1962. The majority of E. adecarboxylata strains studied by Leclerc were isolated from food samples, but studies conducted by the National Institute of Health in Japan discovered that the species is not uncommon among clinical isolates. It is sometimes confused with E. coli because it forms colonies similar to E. coli on general media and shares many phenotypic similarities [5]. Fermentative studies aided in distinguishing the two species, and E. adecarboxylata was assigned to a new genus, Leclercia due to its importance in food microbiology and in honor of the French bacteriologist [5]. Leclerc lived during the forefront of a taxonomic revolution, directly experiencing and impressively assessing the impact 16S RNA would have on classification and identification of bacteria in his own authorized publication, Microbiologie générale [6].
Since then, one more validly published species has been added to the genus, L. pneumoniae. Along with the two valid species, "L. tamurae" was published in 2022 but is still not considered a valid species. Since this proposed species was recently published, it is still awaiting judicial review from the Judicial Commission of the International Committee on Systematics of Prokaryotes (ICSP) in order to become validly published.
How can we identify bacteria at a species level?
Relatively higher homogeneity among the organisms within the Enterobacteriaceae family pose challenges for effective microbial identification at a species level. Among the various commercial methods of microbial identification processes, modern proteotypic and genotypic methods are commonly used. The proteotypic method Matrix-Assisted Laser Desorption/Ionization-Time of Flight (MALDI-TOF) can be used to identify Leclercia adecarboxylata. This proteotypic method provides distinctive spectra for L. adecarboxylata compared to other closely related species. Nonetheless, the process is limited by the strain diversity representing the species in the MALDI database since environmental and growth conditions, along with sample preparation, alter the spectral profile of the organism.
Among the genetic methods of bacterial characterization, the 16S rRNA gene is the most common gene used for microbial identification. However, due to the substantially conserved 16S rDNA, Leclercia adecarboxylata is not distinguishable from Enterobacter cloacae, E. kobei, and E. ludwigii when using this genotypic identification method. As a result, for genotypic identification of this organism at the species level, we would have to use an alternate gene that is less conserved but best represents the species tree. A single conserved housekeeping gene, leuS, was identified among these four closely related species. Phylogenetic analysis based on the leuS gene showed that it could provide species-level identification for the samples that belonged to any of the four species. Furthermore, in the case of the identification of an objectionable organism such as L. adecarboxylata in a production facility, it becomes vital to utilize a multilocus sequence typing approach to determine the contamination source for strain tracking and limiting further outbreaks.
References
- Maude adverse event report: Cerus corporation intercept blood system for platelets. accessdata.fda.gov.(n.d.). Retrieved October20,2022, from https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfmaude/detail.cfm?mdrfoi__id=10292518&pc=PJF
- FDA / CBER Resources Page. Important Information for Blood Establishments and Transfusion Services Regarding Bacterial Contamination of Platelets for Transfusion. U.S. Food and Drug Administration website. Retrieved October 20, 2022, from https://www.fda.gov/vaccines-blood-biologics/safety-availability-biologics/important-information-blood-establishments-and-transfusion-services-regarding-bacterial
- Zayet S, Lang S, Garnier P, Pierron A, Plantin J, Toko L, Royer PY, Villemain M, Klopfenstein T, Gendrin V. Leclercia adecarboxylata as Emerging Pathogen in Human Infections: Clinical Features and Antimicrobial Susceptibility Testing. Pathogens. 2021 Oct 28;10(11):1399. doi: 10.3390/pathogens10111399. PMID: 34832555; PMCID: PMC8619052.
- Leclercia. Leclercia - an overview | ScienceDirect Topics. (n.d.). Retrieved October 20, 2022, from https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/leclercia
- Tamura, Kazumichi, et al. “Leclercia adecarboxylata Gen. Nov., Comb. Nov., Formerly Known Asescherichia adecarboxylata.” Current Microbiology, 1986 13(4): pp. 179–184, https://doi.org/10.1007/bf01568943.
- Leclerc, Henri. Microbiologie Generale. Doin, 1975.
Ryan Cox joined Charles River Laboratories in 2019 after graduating from the University of Delaware, studying chemistry, then working as a Molecular Technologist in the Sequencing Laboratory. He is currently working with Accugenix's Microbial Databases and Technology Transfer team as an associate scientist. He assists in the ongoing curation and maintenance of the MALDI-TOF Library, developing strain typing assays and other various projects to help improve the workflow of the laboratories. In his free time, he enjoys running, cooking, playing Dungeons and Dragons, and spoiling his cat, Butter.




