
Discovery
|
Emanuele Loro, Stuart Ward, Alessia Arduin
Fighting COVID-19 with Novel Hit Compounds
Three of Charles River’s Early Discovery teams join the hunt for new drugs to treat the deadly coronavirus
Faced with a threat to human health unprecedented in most of our lifetimes, the scientific community worldwide has responded rapidly and collaboratively to try to develop prophylactic and therapeutic agents to prevent and treat COVID-19 infections.
Within the Early Discovery division of Charles River, we’ve joined this effort in the hope of contributing what we can to alleviate the suffering experienced by so many in the current pandemic. Three of our teams have been working together using their own specific skills to help identify novel hit compounds that could form the basis for drug discovery programs targeting the virus.
The first component required in a hit discovery toolbox is a biochemical assay in which we can rapidly test compounds of potential interest to see if they show useful activity. In just six weeks, Early Discovery’s biology team developed a viral protease inhibition assay specific to COVID-19. Like other coronavirus proteases, the COVID-19 protease (SARS-CoV-2 or MPro) is responsible for the cleavage of the long string of precursor proteins encoded by the viral genome into individual mature proteins. Such proteins enable new viruses to be packaged and spread within the body and so represent an excellent target for antiviral drug development. With this in mind, we focused on targeting MPro activity. This protease was recombinantly expressed in bacteria and purified in-house by applying and optimizing conditions described in the literature. The assay uses a fluorescent version of a small amino acid sequence that is recognized and cleaved by MPro. We have modified this sequence so that cleavage by MPro results in the emission of a fluorescent signal that we can measure as indication of MPro activity. By performing multiple iterations, we optimized several experimental parameters, e.g., amount of MPro and fluorescent substrate, composition of the reaction mix, time of incubation, until we obtained a robust assay that we can now apply to screen for molecules capable of inhibiting MPro activity.
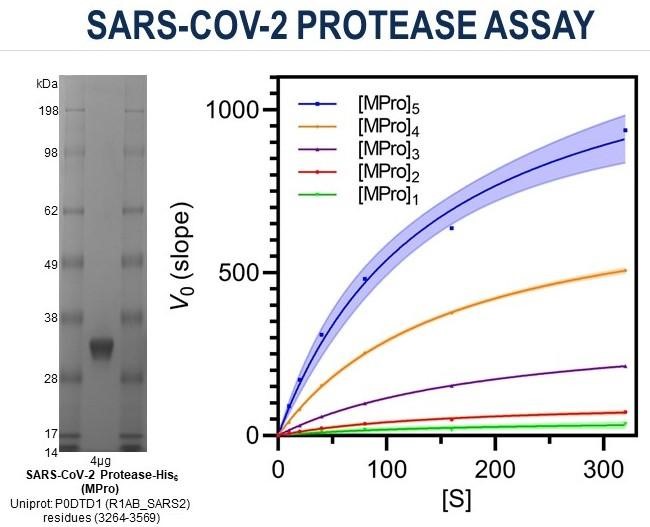
Figure 1: characterization of MPro protein and assay
While the assay was being developed, our teams of chemists and Computer-aided Drug Design (CADD) scientists were already busy. In March 2020, the team at the Diamond Synchrotron (near Oxford, UK) published a new crystal structure of MPro together with the results of a successful fast-tracked crystallographic screen of its XChem fragment collection. Using this information, Charles River’s UK Discovery Chemistry and CADD teams joined the global effort to help combat COVID-19 by providing compound designs for synthetic consideration and ultimately biological testing.
Specifically, we responded to the call to assess the covalently binding fragment hits which have the potential to be highly selective for MPro. A truly collaborative effort between the two departments assessed the binding sites of proximal fragments, designed hybrid molecules, re-assessed them through docking and prioritised a list of 20 compound designs to upload all within one week to meet the deadline for the first round of covalent inhibitor submissions. We are currently waiting to hear which covalent inhibitors have been selected for synthesis, which will hopefully include some from our work.
In parallel to exploiting the fragment screening data, we also assessed commercially available covalent inhibitors of other proteins in the hope that they might provide further starting points for compound design or even have potential themselves for re-purposing as COVID-19 treatments.
Finally, in addition to the work mentioned above, a virtual screening campaign was launched by the CADD group to identify compounds from Charles River’s Diversity Collection, which comprises 610,000 molecules. Using three X-ray structures of MPro, a complex workflow was followed that saw the compounds docked into all three active sites with increasing levels of sophistication. A consensus selection procedure taking account of docking scores and ligand pose strain energies resulted in a set of 3628 compounds being chosen for visual inspection. This time-consuming final step was divided amongst members of the CADD team and led to the ultimate selection of 684 compounds. These are now ready to be plated and screened in the assay mentioned earlier.
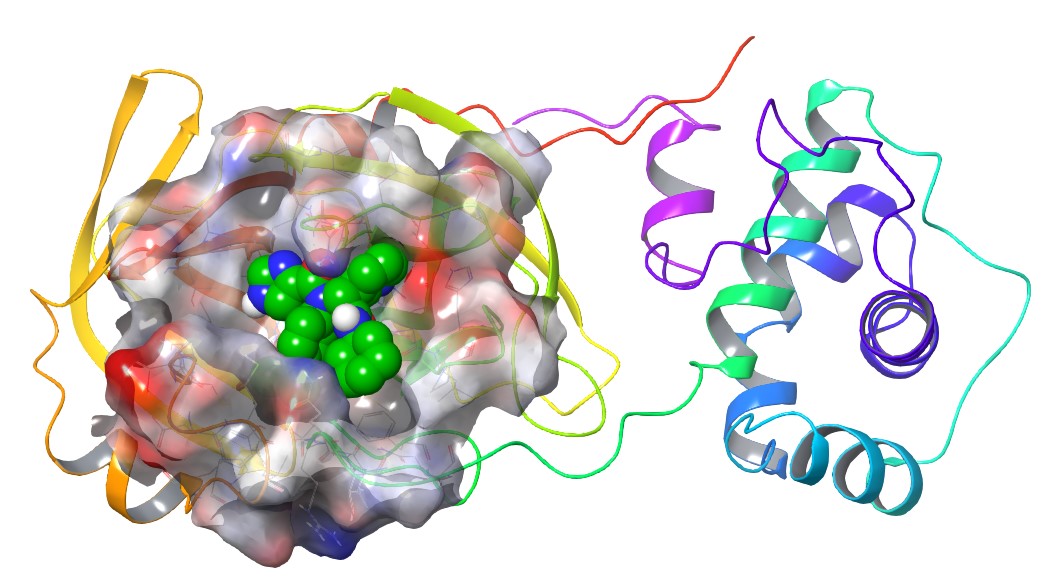
Figure 2: X-ray structure of MPro showing binding of a ligand to the active site (PDB entry 6W63)
In carrying out all the above activities, Early Discovery has illustrated its ability to respond rapidly and flexibly at this critical time to the needs of the scientific community, and indeed the wider world.
The following Early Discovery groups contributed to this work and article:
Assay Development: Alessia Arduin, Emanuele Loro, Vad Lazari, David Whalley
Structural Biology: Rowan Halls-Kass, Pam Dossang, Paul Wan
Chemistry: Paul Winship, Toby Blench, Stuart Ward
CADD: Luca Carlino, Emanuela Gancia, Silvia Bonomo, David Clark, Lampros Milanos
