|
David Clark, PhD
The Secret’s in the Source
How open-source software is transforming scientific research
I did a fair bit of coding in the early part of my career in computer-aided drug design, and I can remember two instances where my limited mathematical ability brought me to a halt. I simply couldn’t understand what was written in a book or journal article and thus convert it into computer code. Even consulting colleagues only gave partial help. It was deeply frustrating as it hindered potentially interesting lines of research. The details aren’t particularly important, although for anyone who’s interested, the two problems were tetrangle inequality bound smoothing and the constructive enumeration of molecular graphs. I know, even the names sound forbidding, so perhaps I shouldn’t be too hard on myself!
These difficulties were encountered early in my career, during the late 1980s and early 1990s – before the internet and World Wide Web were part of our everyday lives. There was very little help available for coders, apart perhaps from the algorithms published in the indispensable reference book “Numerical Recipes” or some pseudocode in a journal article. Fast forward some 35 years to the present day, and things are very different. There are innumerable resources “out there,” and it is far easier to contact scientists worldwide to seek help, advice, and even actual code to solve problems. Nowadays, the two roadblocks I encountered could probably be solved relatively straightforwardly by recourse to source code published freely on the internet for tetrangle inequality bound smoothing and chemical graph generation.
These are instances of what we now call “open-source” software, which Wikipedia defines as “computer software that is released under a license in which the copyright holder grants users the rights to use, study, change, and distribute the software and its source code to anyone and for any purpose”. Allied to the development of the internet has been a remarkable growth in this apparently altruistic sharing of software. Platforms like GitHub and SourceForge host hundreds of thousands of programs and have become “go-to” resources for developers and users of scientific code. This sharing is not only the preserve of individuals; even large organisations make software that they have developed available for all to use in the pursuit of scientific goals.
A prime example of the latter, and one that we have been glad to benefit from in the development of our Design and Predict Hub, is the REINVENT4 program, developed and made open-source by AstraZeneca (AZ). REINVENT4 is a suite of tools for the construction of potentially novel, drug-like chemical structures using the techniques of generative AI. AZ has published an example of how REINVENT4 has been used on one of its drug discovery projects to help in the design of novel Cbl-b inhibitors. The Figure below shows how the LibInvent module of REINVENT4 was employed to design a library of more than 20,000 compounds, which were triaged by further computational methods to just three compounds for synthesis. Medicinal chemists then optimised the initial novel idea (compound 2) to give the final compound 5, which contains the novel moiety (shown in blue) proposed by REINVENT4.
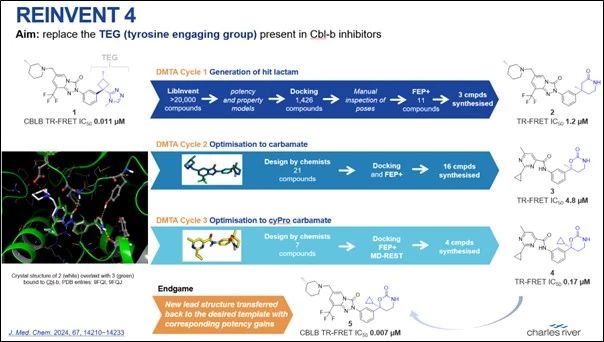
Figure credit: Elisa Pasqua (CRL)
It’s hard to imagine the number of person-hours that must have gone into the development of REINVENT4, and so the fact that it is freely available still seems remarkable to me.
In my field of computer-aided drug design (CADD), there is a vast range of high-quality, open-source tools available, such as RDKit for cheminformatics, AutoDock Vina for docking, and GROMACS for molecular simulation, to name just a few. Such tools are used daily by scientists across industry and academia. Some significant open-source software development projects have been brought together under an umbrella organisation: the Open Molecular Software Foundation, whose stated mission is to “Advance humanity’s ability to understand and shape the physical world through open technologies”. The OMSF team has recently published a substantial Perspective, presenting the organisation and outlining the growing role of open source software in molecular modelling.
At first sight, then, everything looks rosy. But are there any potential pitfalls or downsides to this open-source revolution? Perhaps just a few come to mind.
First, in this era of interconnectedness and sophisticated hacking, we need to be sure that our use of open-source software does not introduce any vulnerabilities into our computing infrastructure. This is a real risk, but fortunately, there is a burgeoning array of companies and tools offering to mitigate the danger.
Second, it’s worth being aware that not all open-source licences are equal. Popular licence types include MIT, BSD 3-clause, Apache 2.0 and GNU General Public. A helpful comparison grid can be found here. It is worth checking carefully (perhaps with the help of a legally qualified colleague if available) to ensure that there are no restrictions that would affect the intended use case.
Third, there is a legitimate concern about the long-term, sustainable support for open-source tools. Many software projects rely on very small developer teams, sometimes just one or two individuals who maintain the software in their spare time. As our reliance on these tools grows, we need to find a way to guarantee long-term support and maintenance of them, or we will be left high-and-dry at some point in the future. The aforementioned OMSF is starting to address these concerns by providing more formal infrastructure and support for some important projects, but many other pieces of software will not fall under its auspices.
Fourth, with so much open-source software available, there is the possibility that time could be wasted evaluating software that, in the end, doesn’t prove useful. An instance of this happening with one particular piece of software has been published. Thus, some self-discipline, reflection and, if possible, discussion with colleagues is required before embarking upon the evaluation and/or exploitation of an open-source solution.
Finally, might it be the case that having everything “off the shelf” might lead to de-skilling and a corrosion of problem-solving abilities amongst software developers? This could be a concern, especially when the rush towards AI-generated code is also factored in. The AI company Anthropic has actually conducted a study on the latter point, from which one of the conclusions was that “Given time constraints and organizational pressures, junior developers or other professionals may rely on AI to complete tasks as fast as possible at the cost of skill development—and notably the ability to debug issues when something goes wrong.”
The above caveats notwithstanding, the astonishing growth in open-source software in recent years has the potential to be a boon for scientific research, including drug discovery and CADD, in which my Charles River colleagues and I are engaged.
I just wish it had happened 35 years earlier …!

