Safety Assessment
|
Brian P. Ruvolo
The Acid Test for Mycoplasma Contamination
Before hopping on the NAT-train, there are several factors to consider
Regulatory guidance mandates that any biological product produced in cell substances (e.g., viral vaccines, monoclonal antibodies, therapeutic proteins, immunological modulators, interferon and other cytokines, growth factors and similar products) must be tested to ensure the absence of mycoplasma. This testing is also required for any cell culture substrates (master and working cell banks, virus seed, each lot of product harvest concentrate and end-point cells) and additives, raw materials and biological implants.
For decades, this had been performed by following the traditional broth/agar culture methods and the indicator cell culture methods, as described in Section 21 of the Code of Federal Regulations (21 CFR 610.30, which did not mention an indicator cell culture method), the United States Food and Drug Administration “Points to Consider” (US FDA PTC, 1993), the United States Pharmacopeia (USP), the European Pharmacopoeia (EP) and the Japanese Pharmacopoeia (JP). As mentioned in 21 CFR 610.30 and US FDA PTC, mycoplasmal contamination testing must be performed by both the agar and broth media procedure (21 CFR 610.30 and US FDA PTC) and indicator cell culture procedure (US FDA PTC), or by a procedure demonstrated to be comparable. Though polymerase chain reaction (PCR) assays have been used for over 20 years as a way in which to detect mycoplasmal contamination, in 2007 the EP outlined an alternative method (Supplement 5.8) where test material could be screened for the presence of mycoplasma by a nucleic acid amplification technique (NAT)-based method.
A Faster Method
NAT-based methods are designed to amplify nucleic acids extracted from a test sample with specific primers that reveal the presence of a particular nucleic acid. Since the traditional mycoplasma assay is commonly the rate-limiting process in the release of a product (28 days), there has been a significant industry-driven push for the approval of a much quicker NAT-based method. This drive has led to numerous companies scrambling to develop their own rapid microbiology method (RMM) via NAT-based assays for the detection of mycoplasma or even develop and market their own manufactured NAT-based mycoplasma detection kits.
Developing an assay based on a NAT platform means one can potentially screen a sample aliquot for mycoplasma and receive results in a matter of hours, completely altering the turnaround time paradigm. Before getting on board the NAT train, however, there are several factors to consider.
Currently, no single NAT-based method or manufactured kit has received regulatory approval for use as a broad-range method of mycoplasma detection. Just as it is required to execute a complete and thorough validation of the traditional method for use in the evaluation of a drug product for the presence of mycoplasma, so too is the same when deciding to use a NAT-based method. As NAT-based methods tend to be much more complex and involved, the standards used to validate these new methods are even more extensive and rigorous than those for the traditional methods.
Secondly, the type of material being evaluated can alter the assay’s sensitivity and ability to detect the presence of mycoplasmal nucleic acids. Samples consisting of a high concentration of cells or cellular debris can interfere with the nucleic acid extraction procedure, effectively “overloading” the isolation system, reducing overall yield of mycoplasmal nucleic acids. In addition, with more non-mycoplasmal nucleic acids present, cross reactivity is always a concern. Depending on the specificity of the primer/probe set and the contaminating nucleic acids present, a false positive is possible. Though some NAT-based kits are designed to better handle high-density cell samples than others, an increase in assay interference and decrease in assay performance due to sample matrix should always be a concern.
Thirdly, one of the most important parameters that should be investigated is the coefficient of viability (GC/CFU) ratio of your mycoplasma control strains. In the most simple of terms, this ratio compares the total number of viable cells that, once inoculated out across the surface of an agar plate, form colonies, versus the total number of copies of the target sequence present. The greater that GC to CFU ratio (5:1 versus 10:1, where the most ideal would be an equal 1:1 ratio), the greater the overestimation of the limit of detection of the NAT-based method employed.
Finally, these methods are based on the detection of amplified nucleic acids, rather than a method which can directly detect intact viable mycoplasma. This has raised concerns that the NAT-based methods and manufactured kits, which claim sample to results in mere hours, may be unable to accurately delineate between living and dead mycoplasmas and free nucleic acids. In addition, if your reaction reagents have not been verified to be nucleic acid-free, there exists the potential that one of your reagents can lead to a positive signal.
The Burden of Proof
Regardless of the source, any positive signal will call into question the safety of the sample, thus putting the burden of proof that the sample is free of living mycoplasma on the shoulders of the end-user; it becomes significantly difficult to assure a lot of material is safe when initially associated with a positive result. You could follow up any NAT-based positive result with the traditional method, of course, but this would only add to the initial turnaround time, leading to a longer timeline than had the traditional method been executed initially. There is also no guarantee that this approach would be accepted by the regulatory agency.
Face it. Would you trust the timely release of your billion-dollar drug product on an assay that has the propensity to yield a positive result, not necessarily due to the presence of viable mycoplasma in the sample, knowing that the result may lead to the destruction of an entire lot of that marketed product? I do not know of many pharmaceutical companies that would be willing to accept that level of risk.
Still, there are many advantages to a NAT-based assay. NAT-based technologies have been around for years and most institutions should have the equipment readily available. Moreover, the quicker turnaround time allows pharmaceutical companies to get their drug products to the market faster. In addition, if this approach becomes more widely used, the costs associated with the execution of an assay will be less and less, which is more cost-effective than executing a study based on the traditional method. Finally, as this technology advances, sensitivity may increase, specificity may become more accurate, and delineation between viable and nonviable mycoplasmas and free nucleic acids may become possible.
Until then, however, it is important that every end-user weigh the risks associated with less-robust NAT-based methods and other RMMs, taking into consideration return on investment, operating income, potential for recall or costs associated with the destruction of a perfectly fine lot of product.
Agar plates inoculated with Mycoplasma orale (left) and Mycoplasma pneumoniae: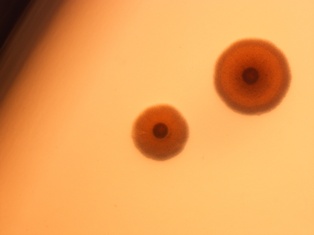

Learn more about Charles River's full Mycoplasma Testing Capabilities
