Discovery
|
Rana Samadfam, PhD
Neglected Players in Microbiome Research
The complex synergies between naturally-occurring viral phages and their bacterial hosts
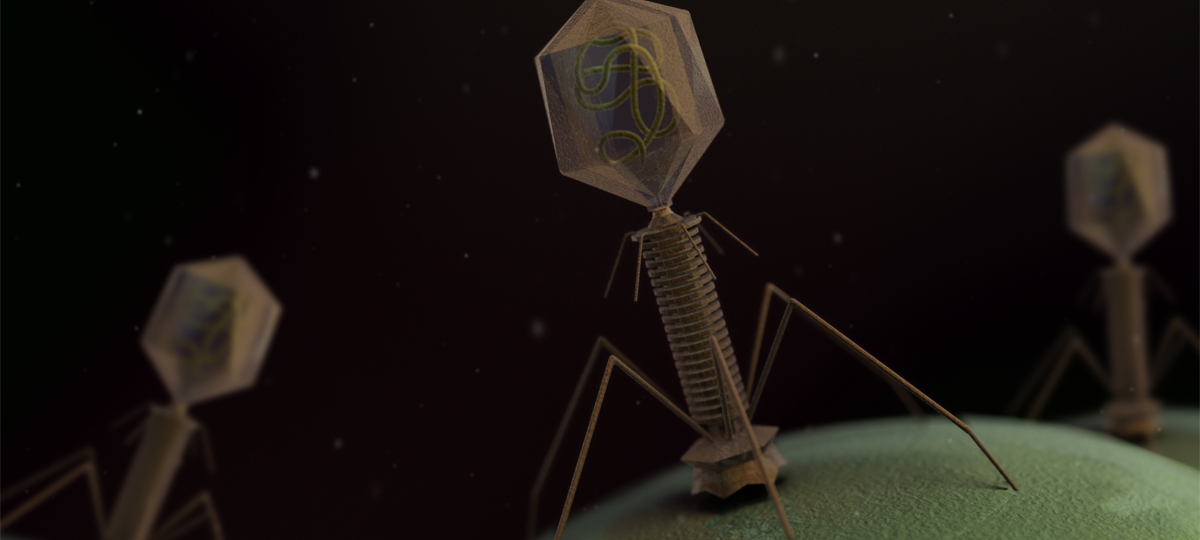
When microbiome research gained the attention of scientists, most of their efforts focused on bacterial populations. They neglected other microorganisms, such as phages and viruses. With renewed attention on phage therapy and advances in gene editing technologies, more research is directed towards the sophisticated interactions between phages and their bacterial host.
Phages have two types of viral reproduction: the lytic cycle and lysogenic cycle, also known as prophage. In the lytic cycle, the phage infects the host and dismantles its genome; hijacking transcription machinery to use for its own replication. Once the host cell is exhausted and cannot further support  replication processes, the newly formed phages will destroy the host cell and each will begin a journey to find and infect a new host. During the lysogenic cycle, the genetic material of a phage is inserted into the genome of the bacteria. During this cycle the bacteria continues to function normally, growing and replicating for generations, with all offspring silently carrying prophage.
replication processes, the newly formed phages will destroy the host cell and each will begin a journey to find and infect a new host. During the lysogenic cycle, the genetic material of a phage is inserted into the genome of the bacteria. During this cycle the bacteria continues to function normally, growing and replicating for generations, with all offspring silently carrying prophage.
A prophage can enter the lytic cycle upon activation by different factors, including stress or inflammatory stimulation. Phages used to combat infection are obligatory lytic phages with limited ability for insertion within the host’s DNA. In a healthy ecosystem (including the human gut), there is a balance between lytic phages and prophage. Several regulatory phage proteins and their expression govern the different phage cycles, and subsequently the fate of the infected host. Mutations in these regulatory genes could influence the ability of phages to destroy the infected cell and its membrane.
The viral-bacteria dance
Transduction is also important in the evolution of microbiome ecosystem. The transduction is the transfer of genetic material from one organism (such as bacterium) to another by viruses including phages. Newly introduced genetic material has potential to modify function of the host or its response to the bacteriophages. Transduction could occur when prophage is activated and the phage genome is excised from the host DNA. During this process, the phages may take away part of the bacterial DNA with them. Transduction may also occur in lytic cycle by incorporating dismantled bacterial genetic material into a new virion during the packaging process, where the components of the virus are assembled like a car on an assembly line.
Mutations and transductions in both phage and bacteria are part of a system called antagonistic coevolution (AC) that amounts to a kind of biological arms race, usually between a parasite and a host. Microbiologist Pauline D. Scanlan used AC in the context of microbiome research to refer to the reciprocal evolution of bacterial defense systems versus the phage’s infectivity. AC between bacteria and bacteriophages is essential for maintaining microbiome diversity by determining which microbial communities will emerge as the strongest and fittest.
The concept of AC also applies to humans and their parasites, like bacteria and viruses. In a human ecosystem, the individual’s health status and diversity of their microbiome are influenced by both evolutionary process and external factors such as diet, hygiene, and exercise (see image below).
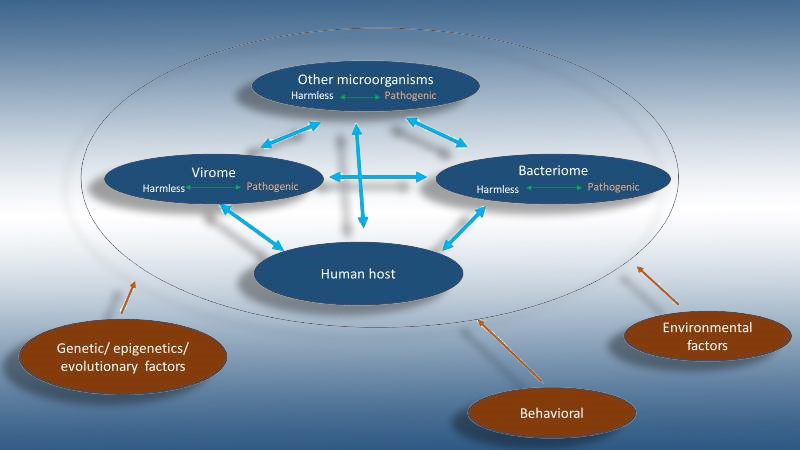
Microbiome research used to be largely dedicated to the bacterial component of the microbiome, with limited research on phages and other players in this ecosystem. Furthermore, researchers have mainly focused on the external factors, with less attention paid to evolutionary internal factors. These static views were necessary first steps in understanding the enormous complexity of the ecosystem and the interactions between different players.
Deciphering our ecosystem
Microbiome research is still in its infancy. Much like solving a puzzle, we must break the ecosystem into different pieces and investigate individual interactions in order to obtain a more comprehensive view of different players and the rules governing their interactions. Feeding this information into mathematical modeling will be the next step toward improving our understanding of our dynamic ecosystem. Mathematical models can be used to simulate microbial community dynamics, such as exploring how a host cell and phage virus impact viral evolution. By widening our knowledge, approaches to treating microbial imbalances will also evolve, and hopefully will include a dynamic and multi-directional approach.
Our next installment is in the phage therapy series will look at the microbiome (including the phage) and its impact on irritable bowel disease.