What You’ll Find in the Complimentary Charles River Cancer Model Database for Preclinical Oncology Research
The Cancer Model Database gives you access to comprehensive molecular data, generated for our PDX models, as well as for CDX and cell lines, to help you find the best-suited models for your preclinical goals.
Our integrated approach using in vitro 2D, ex vivo 3D, and in vivo models, as well as bioinformatics data, facilitates drug development and accelerates your oncology research. You can search by:
- Tumor metadata including patient data, growth characteristics, and histology
- mRNA expression level
- Whole exome sequencing mutation profiles
- Gene copy number alteration
- And more
Search criteria can be used individually and in combination.
Thanks to this Charles River Database, there are no more animal model errors, and it helps us refine our studies.”
Academic Researcher
How to navigate efficiently through our Cancer Model Database
Search by:
- Model type, such as PDX, CDX, and cell line
- Cancer type
- Gene expression, mutation, and/or copy number variation
- Predicted HLA types
- Tumor metadata (such as histology, patient characteristics, etc.)
Dive Deep into our Open Access Cancer Model Database
Our cancer model database has different types of molecular data:
- Gene expression determined by Affymetrix® Human Genome U133 Plus 2.0 Array
- RNA-seq data (gene expression, mRNA mutations, fusions, HLA predictions)
- Gene mutations determined by whole exome sequencing
- Gene copy number variations determined by Affymetrix® Genome-Wide Human SNP Array 6.0
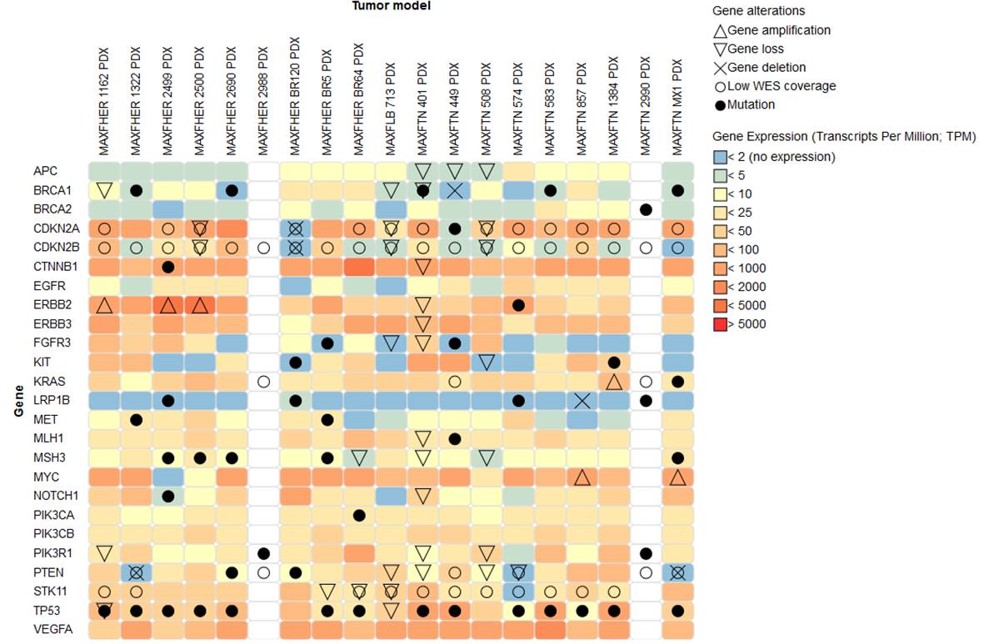
You can easily visualize your search and download the data for further analysis. This image is a visualization example including amplification, loss, deletion, mutation, and expression of the gene of interest across different models.
Your search results indicate for which assay type the model(s) are available: 2D screening, 3D ex vivo and/or in vivo.
Our experts are ready to answer your questions.
Oncology Preclinical Study Planning Toolkit
Take control of your efficacy study by getting instant access to study pricing, cancer model data, and connection to our oncology specialists, so that you feel supported on your oncology efficacy journey.
Find the Right Model
In vivo: CDX and PDX Model Database
Our cancer model data and pre-selection process ensures you have selected the most relevant model for full-scale in vivo efficacy studies. The Cancer Model Database includes tumor models from a wide range of tumor subtypes, including patient-derived xenografts (PDX), cell line-derived xenografts (CDX), and mouse syngeneic models. You can view our models across a range of different histotypes that closely mimic the respective clinical disease. The Cancer Model Database shows our full range of in vivo models.
Browse the CDX and PDX Model Database
Have a look at the in vitro, 3D, and cell lines database
The industry standard for early efficacy and tumor type identification, in vitro screening is a critical step to identify innovative compounds in a rapid, reliable, and cost-effective way. Simplifying and accelerating what can be an otherwise time-consuming process, our Cancer Model Database is a valuable resource for oncology researchers as they enter the critical in vitro stage of their work.
Using data from the Cancer Model Database, you can select the best in vitro model to test your compound from our selection of 2D in vitro assays and 3D ex vivo assays.
In addition to the Cancer Model Database, Charles River has a range of pediatric cancer models. Since the RACE-Act, all new oncology therapeutics must be evaluated for safety and efficacy in pediatric cancers if the treatment is directed at a molecular target relevant to pediatric cancer, including therapeutics with an orphan drug designation. Find more than 200 pediatric well-characterized PDX models, as well as CDX models available for your research project.
The Charles River’s Cancer Model Database is even more useful than one of the biggest publication databases!”
Academic Researcher
Frequently Asked Questions (FAQs) About the Cancer Model Database
-
What is the difference in gene expression analyzed by Affymetrix® versus RNA-seq?
Unlike arrays, RNA-seq technology does not require species- or transcript-specific probes. Both technologies detect the RNA expression level of a specific gene.
-
How many models express my target of interest?
When searching for your gene of interest all models will be displayed. You can create a cut-off for expression in the results.
-
Can I search by histology?
Yes, you can type in a histology in the search box at the top of the Cancer Model Database.
-
How can I find the mouse strain?
For the syngeneic lines it is annotated in the model as ethnicity/strain.
-
How can I find pediatric models?
The majority of pediatric models are not in the database. Instead, they are available through our collaboration with the ITCCP4 consortium. Learn more on our pediatric cancer drug development page.