Discovery
|
Christoph Eberle, PhD
Profiling Immune Subsets of Syngeneic Mouse Models
Knowing or not knowing which immune cells infiltrate a tumor can help in drug outcomes
There is no doubt that immunology has opened the door to an arsenal of new therapies. Included in this war chest are immune checkpoint inhibitors, which work by blocking an off-switch tumors activate to prevent T-cells and macrophages from attacking them.
Yet while immunotherapies can shrink and, in some cases, destroy tumors, they only work in a relatively small percentage of patients. Why is that so? One reason is that tumors exploit their habitat to survive, seizing on any number of cells and proteins that live in and around it to spook the immune responses that are so important to the outcome of the patient. This so-called tumor microenvironment (TME) has become a kind of playbook for researchers, offering insights into a tumor’s progression, potential response or resistance to treatment.
So how do you know which immune cells will invade a patient’s tumor, and which ones will not?
What the tumor is embedded in depends a lot on the type of tumor. Part of a tumor’s habitat are filled with blood vessels, stroma, endothelial cells, and a variety of immune cells known as tumor infiltrating leukocytes (TIL) whose composition may change through immunotherapy. Ideally, then, you want to recruit the right set of immune cells to clear a tumor.
We can study tumor behavior in vitro, but animal models are still the backbone of biomedical research experiments to look at the interaction between cancer and the immune system. For oncology drug development, murine cancer cell lines that can grow in syngeneic strains are a long-established choice. Those strains retain the ability to mobilize all immune cell combatants, and therefore if implanted with a cell-derived xenograft allow for assessing how a competent immune system fights off mouse tumor burden.
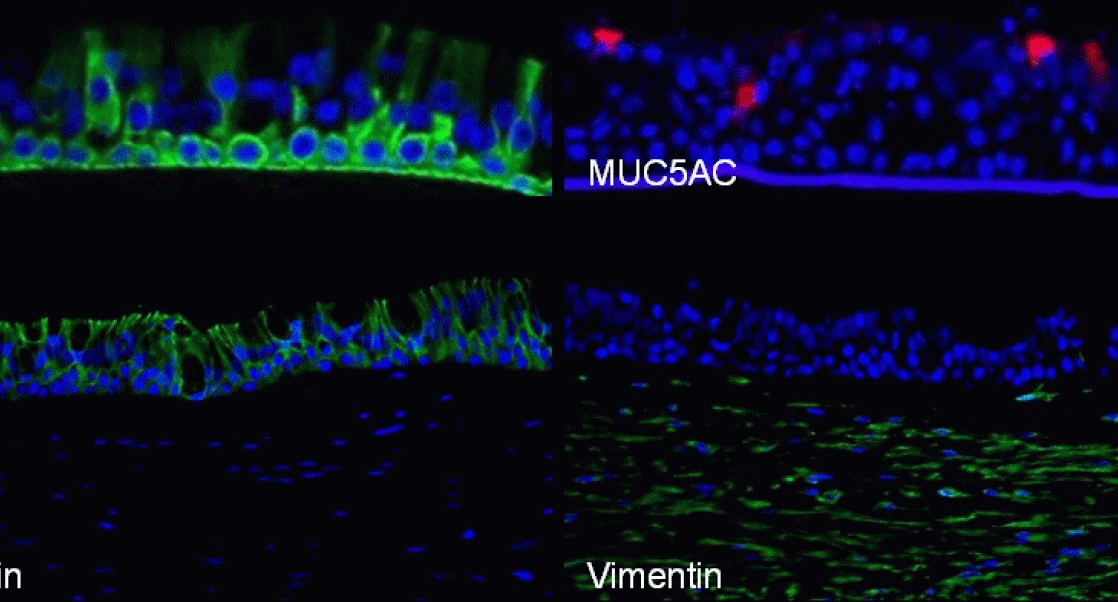
In Balb/c and C57BL/6 mice bred at Charles River, we characterized four models of solid cancer types: colon (MC38) and renal (RENCA) adenocarcinoma, melanoma (B16F10), and mammary carcinoma (EMT6). We compared growth kinetics and profiled the immune cell composition in each tumor type using acoustic-assisted flow cytometry. These analyses confirmed that common lymphocyte subsets of both lymphoid and myeloid origin naturally infiltrate all these different tumor microenvironments.
This kind of profiling is important, if we want to correctly determine if a cancer drug can have a future. Flow cytometric readouts are information-rich, and when considered for early decision-making in drug pipelines, tumor model datasets need to be reproducible. One way to ensure the reliability of these datasets is through TIL immunophenotyping, a tool that helps us to gauge the presence of immune cell populations and their frequency.
TILs are a relevant pharmacodynamic endpoint to evaluate the efficacy of immunotherapies. For this reason, it is critical to characterize various challenging mouse models to know which immune cells coexist in the tumor microenvironment without therapeutic intervention. It resembles scouting out a battlefield, on which some of these cells can be our allies or our foes. We need to able to tell one from the other. This more strategic approach will help us achieve our ultimate mission—better drug responses and better outcomes.