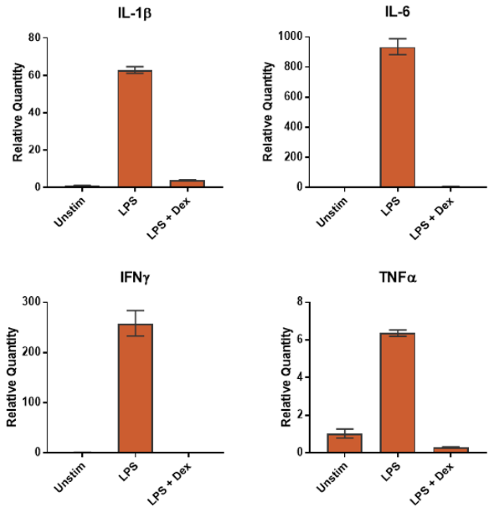
Measuring Gene Expression Using ddPCR and qPCR Services
qPCR was considered the gold standard for measuring changes in gene expression and can detect as little as a single target molecule per reaction. Characterization lends itself well to scenarios where there are either a small number of targets to measure but large numbers of samples or the reverse scenario. ddPCR is the latest and most sensitive approach for companies developing cell and gene therapies, biomarkers, and/or oligonucleotides.
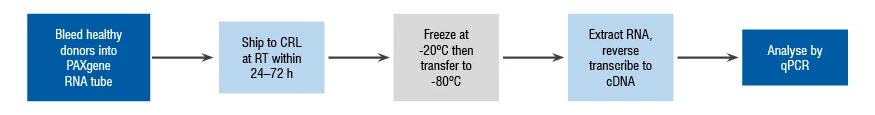
Before analyzing clinical samples, the first step is to identify a suitable sample handling protocol that enables samples to arrive at the clinical laboratory from the clinical site intact and with an appropriate quality level. A representative selection of samples should be subjected to the same storage and shipping conditions that will be used in the clinical trial to validate the pre-analytical sample handling process.
eGuide: Biomarker Solutions from Discovery to Clinical Trials
Learn how strategic biomarker integration can accelerate research & de-risk pipelines to bring life-saving therapies to patients faster. This guide helps you uncover smarter ways to de-risk studies and accelerate timelines, without compromising scientific rigor.
Read the Guide
ddPCR Services and Evaluations
Our experts have fully validated, the largest commercial fleet of QX One instruments for full ddPCR services and evaluations to ensure the most sensitive, precise techniques for our clients developing cell and gene therapies, oligonucleotides, or biomarkers. The detection of ddPCR uses the same chemistries as real-time PCR; however, it allows for more sensitive readouts for your studies. Each partition is individually called as positive or negative based on fluorescent signal resulting in higher quality data and a decrease in variability.
ddPCR Assay Capabilities
- 7 Bio-Rad QX One instruments validated
- Physical partition of reaction
- Oil emulsion or droplets (Droplet Digital)
- Physical portioning (Digital)
- End point PCR method
- Detection using same chemistries as Real Time PCR
- Each partition is individually called as positive or negative based on fluorescent signal
Webinar: The Critical Role of Bioanalysis in AAV Therapies
This webinar will explore the expanding role of bioanalysis in AAV therapies, focusing on biodistribution, transgene expression, immunogenicity, and ELISpot.
Watch Now
ddPCR Bio-Rad QX One Read Out Examples
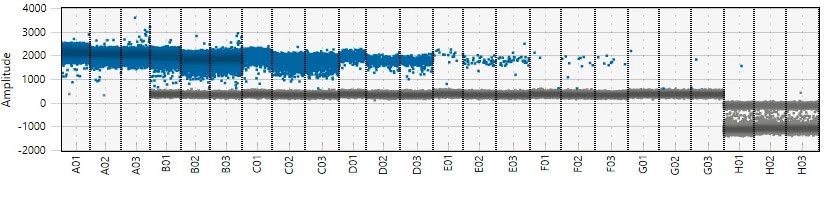
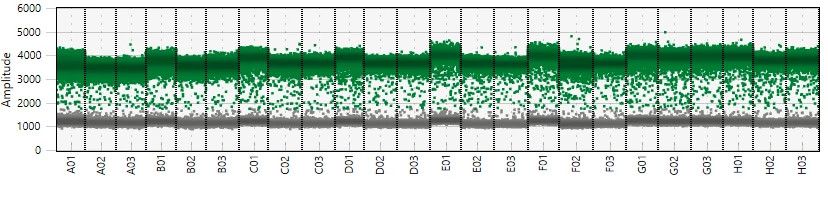
qPCR Assay Validation
While qPCR assays are a well-established technique to measure gene expression, there are factors to carefully consider when validating the method in order to ensure the generation of reliable data.
-
Selection of Reference Genes
The most common approach for gene expression studies is a relative quantitation approach. When implemented, it is crucial that the expression of the reference genes used for normalization are not affected by the therapeutic in order to prevent skewed data. Selection of stable reference genes can be achieved by screening a panel of candidate reference genes in a representative selection of treated versus untreated samples using a reference gene selection algorithm.
-
PCR Efficiency
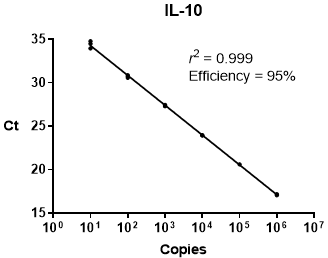
It is important to account for differences in amplification efficiencies of the genes of interest. In a perfectly efficient qPCR assay, the amount of PCR product exactly doubles with each subsequent amplification cycle; however, most assays/primer sets are not exactly 100% efficient. These deviations from 100% efficiency can skew the data if not accounted for in calculations of relative quantities. Once the efficiency of each assay is known, the value is incorporated into the analysis for a more accurate measurement of gene expression.
qPCR Efficiency
-
Limits of Detection and Quantitation
It is important to understand the limits of the assay in terms of lower limit of detection (LLOD) and lower limit of quantification (LLOQ). LLOQ is the lowest concentration of a target that can be accurately quantified, and the LLOD is the lowest concentration of a target that can be reliably detected, but not accurately quantified. Defining these limits is especially important if the targets are expected to be low in abundance. This must be determined experimentally in each laboratory using the same reagents, equipment, and operators that will be used in the analysis of the clinical samples.
-
PCR Inhibitors
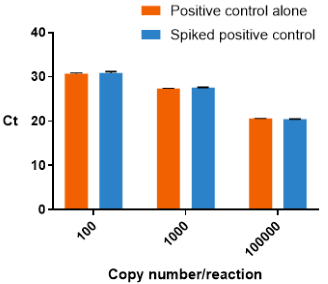
Certain sources of RNA contain significant levels of PCR inhibitors, which can affect qPCR assay performance if they are not efficiently removed during the extraction process. One way of determining whether PCR inhibitors are present in a sample is to take a synthetic DNA template and compare the CT values when measured alone and spiked into a comparable biological matrix. If a significant level of inhibition is observed, the RNA extraction protocol may have to be further optimized to remove contaminating PCR inhibitors.
Testing for the presence of qPCR inhibitors
-
Normal Intra-Donor Variation
Other confounding factors that could be assessed during the validation phase include the normal level of variation within each donor over time. To assess this, samples are taken from donors at several timepoints that are similar to the sampling timepoints that will be used in the trial. A variation of twofold up or down can be considered within normal levels of variation, a threshold that a candidate therapeutic would have to exceed in either direction for the change to be considered potentially significant.
RNA Analysis by qPCR
Do you need guidance on your ddPCR studies?
Frequently Asked Questions (FAQs) About qPCR Services and Gene Expression Profiling
-
What sample types can be analyzed using qPCR assays?
qPCR can be used to analyze many different biological sample types, including cultured cells, whole blood, PBMCs, tissues, fixed-formalin paraffin embedded (FFPE) samples, and cell-free samples such as cell culture supernatants, plasma and cerebrospinal fluid (CSF) to analyze cell-free, or circulating free DNA (cfDNA).
-
How sensitive is qPCR?
For a well-optimized assay, qPCR can detect as little as one copy of target sequence per reaction. For accurate quantitative analysis, the limit is usually around 10 copies per reaction. In terms of fold-changes in gene expression, differences as small as half-fold can be detected.
-
Is multiplexing possible with qPCR services?
Between three and five targets can be analyzed in parallel, depending on the qPCR machine. Additional validation is usually recommended to ensure that multiplexing does not affect the performance of each individual assay.