Measuring Gait Changes in Movement Disorder Models
Impairment of fine motor skills is used as an early diagnosis marker in muscular degenerative and neurodegenerative diseases including Parkinson’s disease. In order to effectively test therapeutic compounds in animal models, it is imperative to characterize the relevant neurodegenerative and muscular degenerative mouse models for fine motor impairment. However, there are limited instruments and assays available to test animal models for fine motor impairment.
Fine Motor Kinematic Analysis Resources
Check out our Fine Motor Kinematic Analysis posters, publications and other scientific resources…
Browse The Resources
At Charles River, we use a novel automated and high-precision system, the MotoRater, to successfully analyze fine motor kinematics in mouse models of movement disorders including Huntington’s disease, multiple sclerosis, ALS studies, Duchenne muscular dystrophy, Parkinson’s disease studies, Batten disease and other neurological and rare diseases. Fine motor kinematic analysis is a highly translational tool as it measures similar parameters in rodents compared to human patients.
Fine motor kinematic analysis reveals early changes in muscular or neuronal function in disease models to support the development of therapeutics that reverse or slow progressive degeneration and to test compounds that can cause fine motor impairment as a side effect. Any rodent model of disease can be analyzed in this system and studies can be customized to measure gait changes while walking, running, wading and climbing.

Neuroscience Translational Tools: Fine Motor Kinematic Analysis
Fine motor kinematic analysis can investigate subtle, yet highly significant, changes of gait and movement details in rodents. Its sensitivity provides a wide therapeutic window for possible treatment candidates, and also enables assessing pharmacological disruption of motor function.
Download the Tech Sheet
Kinematic Data Analysis
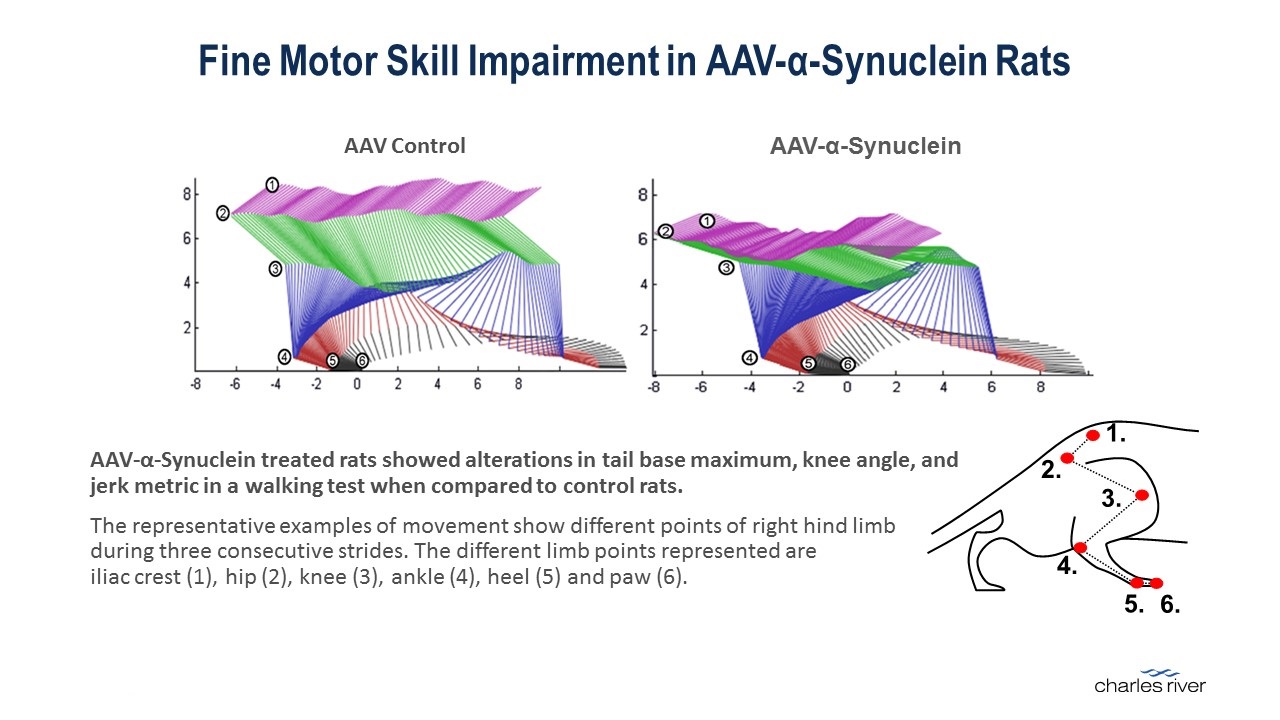
The output from the MotoRater system is high speed video that is analyzed using a proprietary and customized data analysis algorithm that analyzes multiple markers on the animal model to develop detailed quantitative measurement of gait changes that are visualized using composite images and can be further analyzed to develop principal component analysis to identify early stage gait changes and loss of balance. The data output includes over 100 different parameters that are used to study the effect of specific therapies on improving gait changes in animal models of movement disorders.
Kinematic data from a walking test in AAV-α-Synuclein treated rats and control rats.
Frequently Asked Questions (FAQs) About Kinematic Analysis
-
How does the kinematic analysis method compare with the rotarod test?
Both methods are used to assess the efficacy of a therapy on animal motion and balance. However, the rotarod measures one parameter (ie. the time on the moving rod) while the kinematic method measures over 100 parameters of movement including movement at the hip, knee, paw, tail and other limb areas. The parameters are analyzed using proprietary algorithms to generate a complete view of changes in locomotion.
-
What is the output of the kinematic method?
The experimental output is a high-speed movie (300 frames per second) that is analyzed using various tools to generate visual graphs of movement changes and a principal component analysis graph that tabulates changes in select parameters.
-
Can I perform experiments to measure gait changes in my lab?
Yes, in theory. The animal models are filmed on a commercial system (MotoRater) but the crucial data analysis is performed using proprietary algorithms that were built by Charles River data scientists.
-
Can kinematic analysis be performed on rats?
Yes, kinematics analysis can be performed on rats and mice, so there is no species dependence. However, we have predominantly tested mice.